# Exercise 03 Solution {.unnumbered}
# • Solution {.unnumbered}
### Preliminaries {.unnumbered}
Load in dataset and libraries of interest...
```{r}
#| warning: false
#| message: false
library(tidyverse)
f <- "https://raw.githubusercontent.com/difiore/ada-datasets/main/data-wrangling.csv"
d <- read_csv(f, col_names = TRUE)
names(d)
```
## Challenge {.unnumbered}
#### Step 1 {.unnumbered}
```{r}
#1
d$BSD <- d$Body_mass_male_mean/d$Body_mass_female_mean
# 2
d$sex_ratio <- d$AdultMales/d$AdultFemale
# 3
d$DI <- d$DayLength_km/(2 * sqrt(d$HomeRange_km2/pi))
# 4
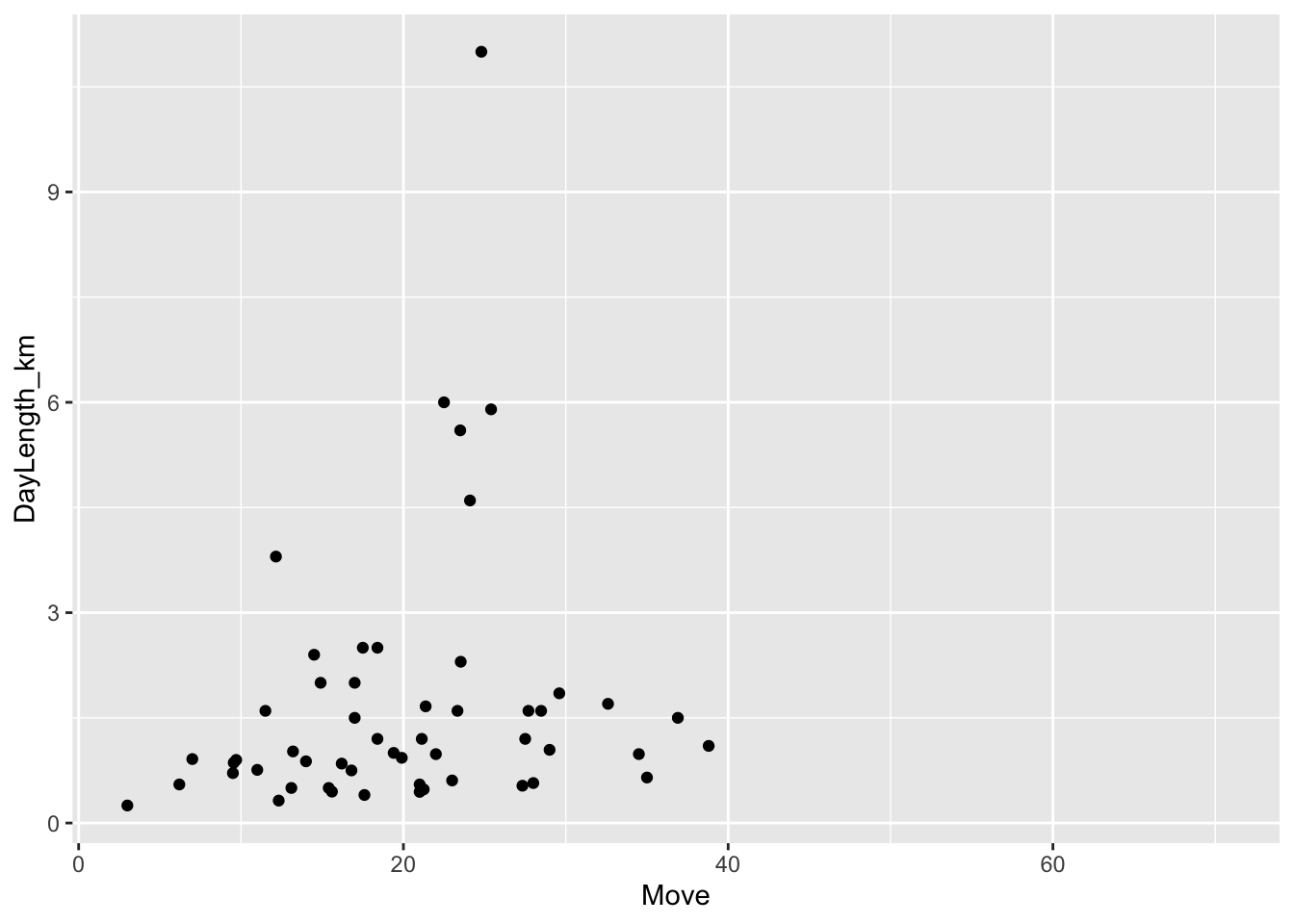
(p <- ggplot(data = d, aes(x = Move, y = DayLength_km)) +
geom_point(na.rm = TRUE))
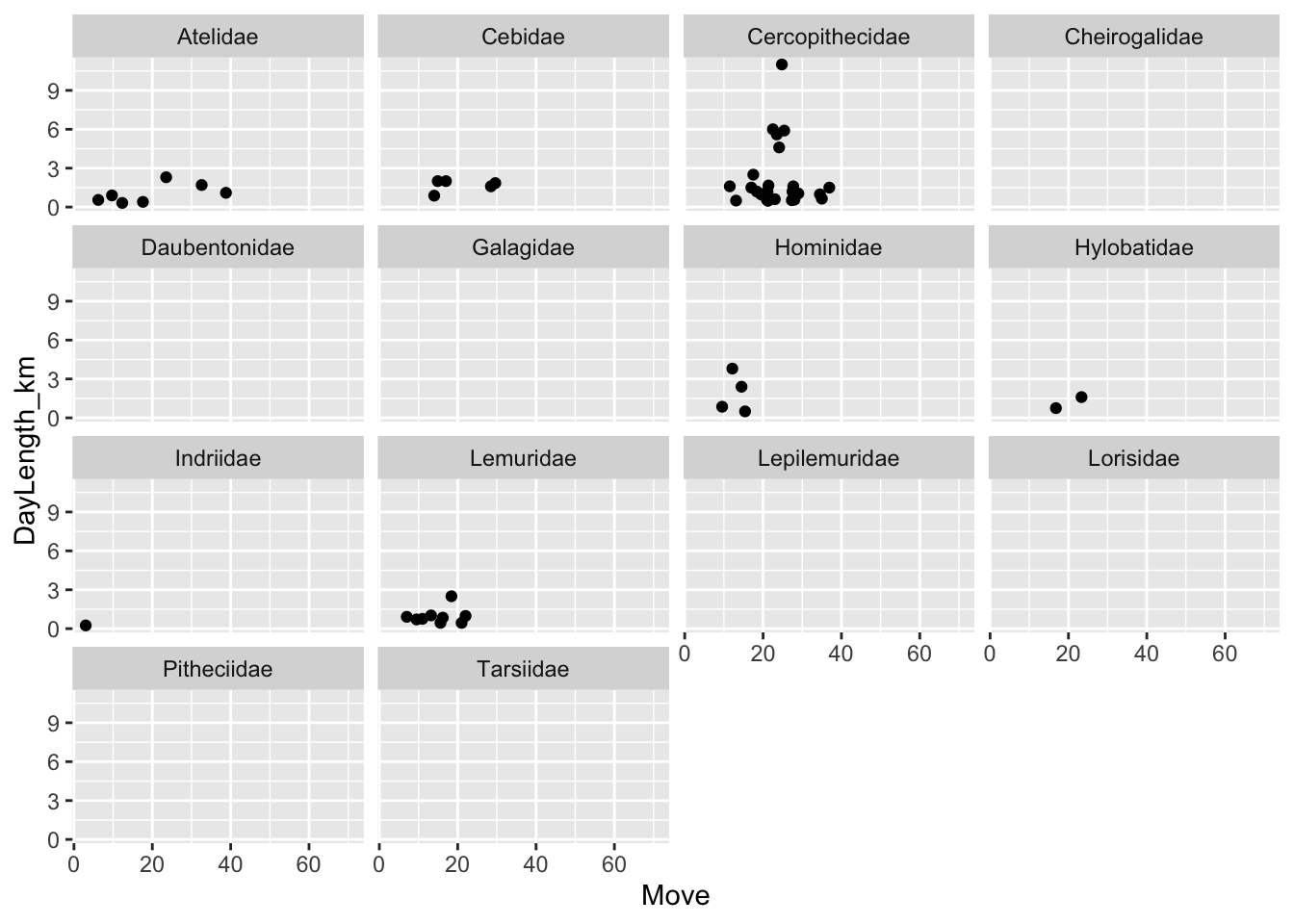
(p <- ggplot(data = d, aes(x = Move, y = DayLength_km)) +
geom_point(na.rm = TRUE) +
facet_wrap(~Family))
# 5
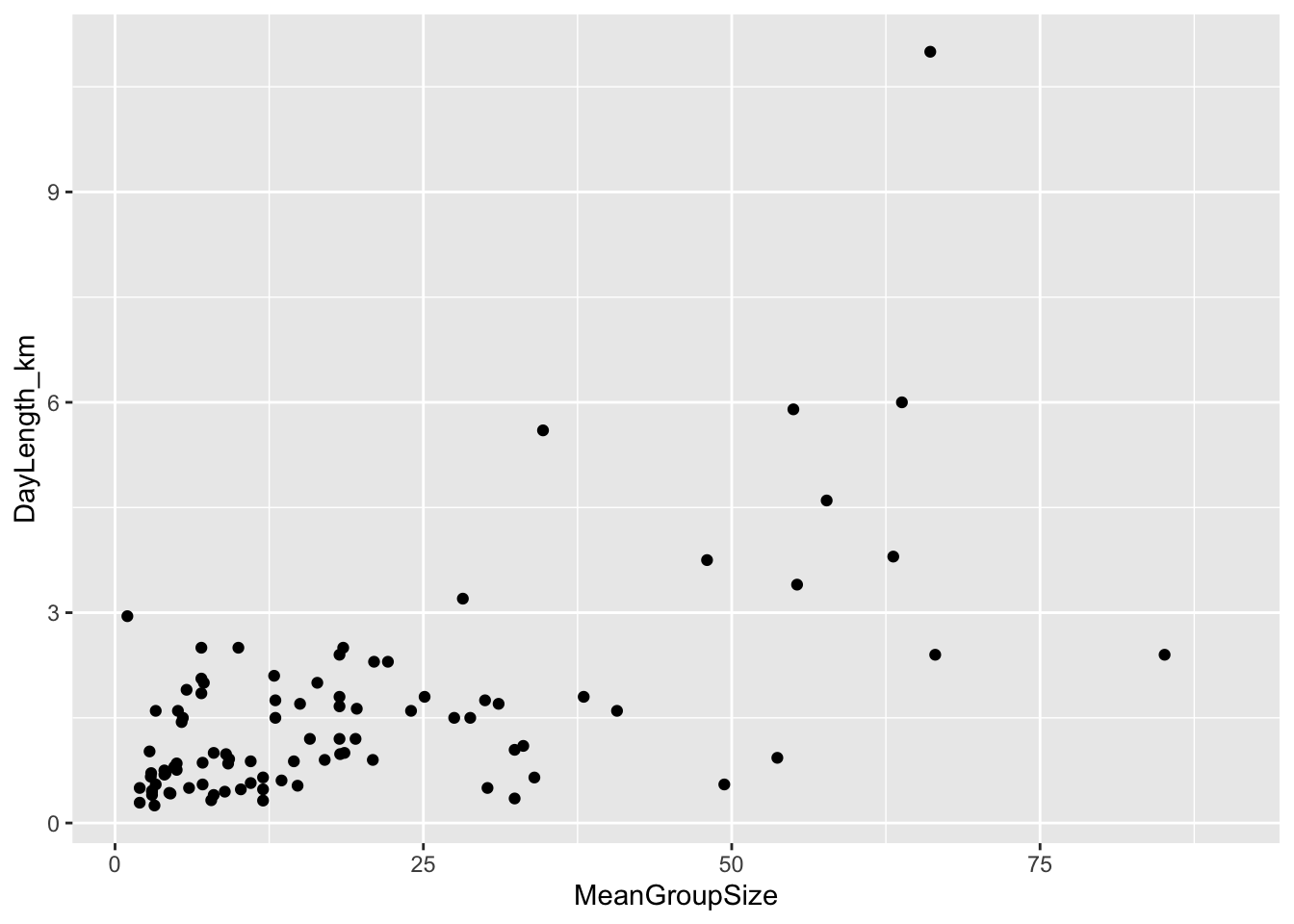
(p <- ggplot(data = d, aes(x = MeanGroupSize, y = DayLength_km)) +
geom_point(na.rm = TRUE))
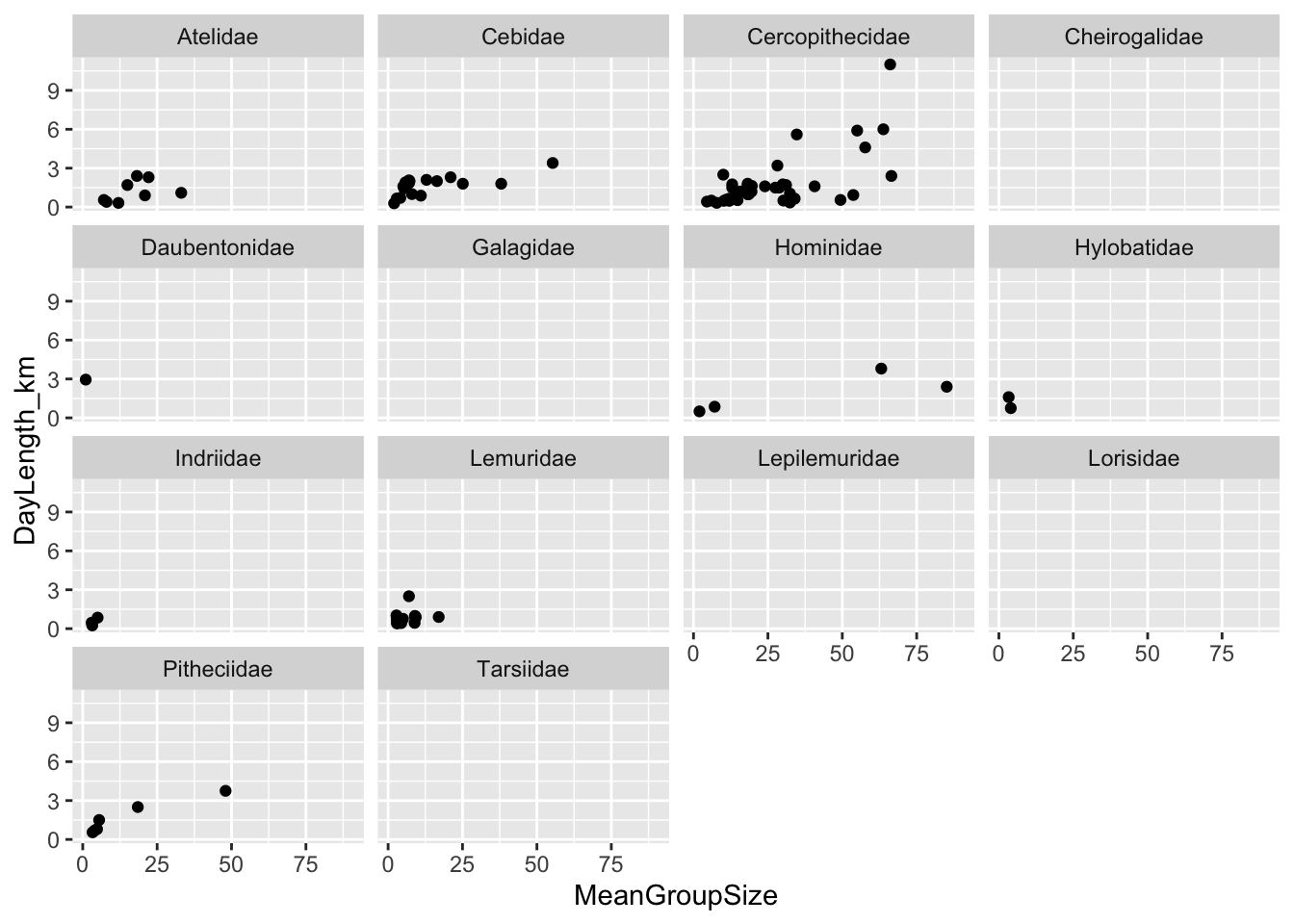
(p <- ggplot(data = d, aes(x = MeanGroupSize, y = DayLength_km)) +
geom_point(na.rm = TRUE) +
facet_wrap(~Family))
# 6
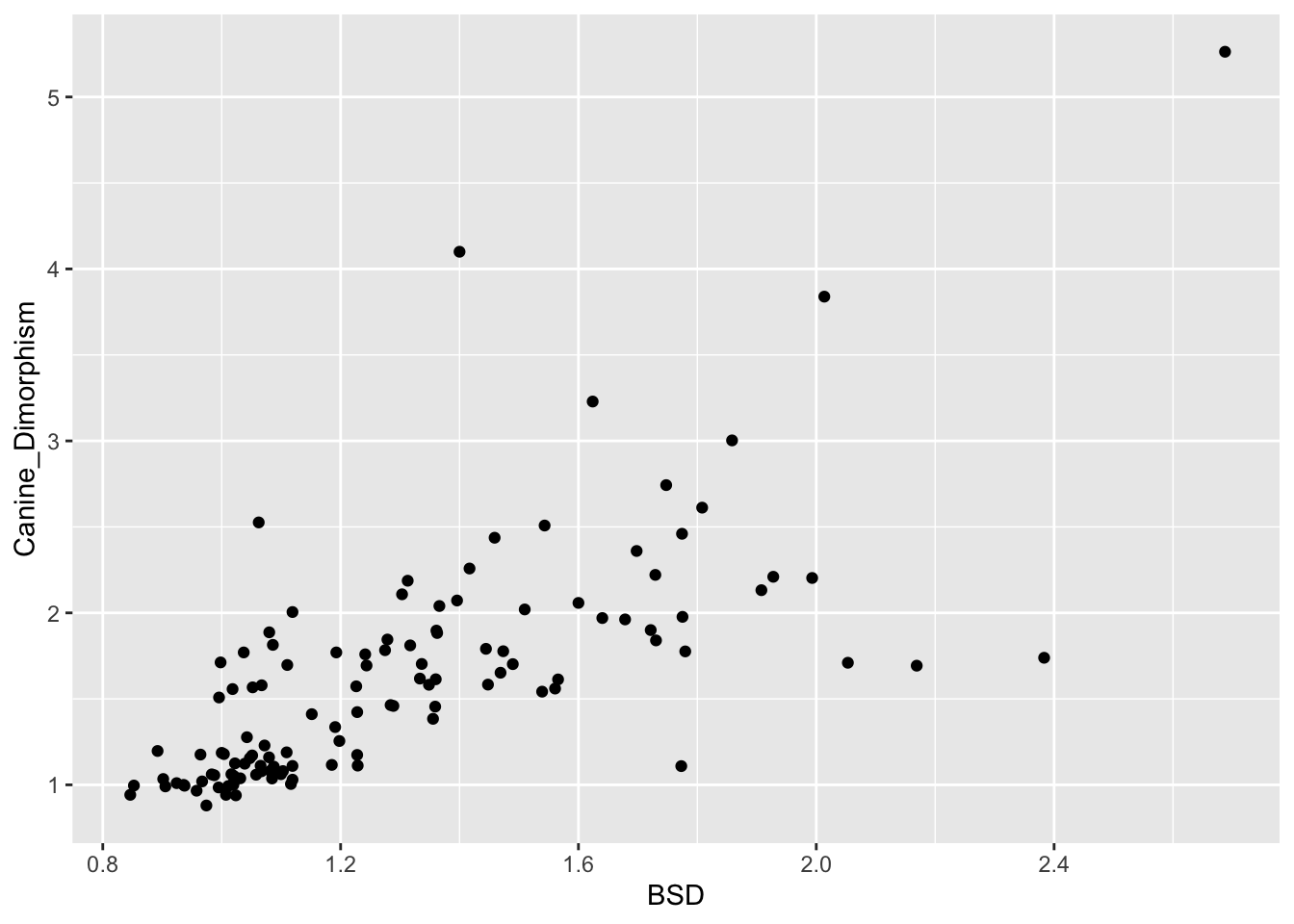
(p <- ggplot(data = d, aes(x = BSD, y = Canine_Dimorphism)) +
geom_point(na.rm = TRUE))
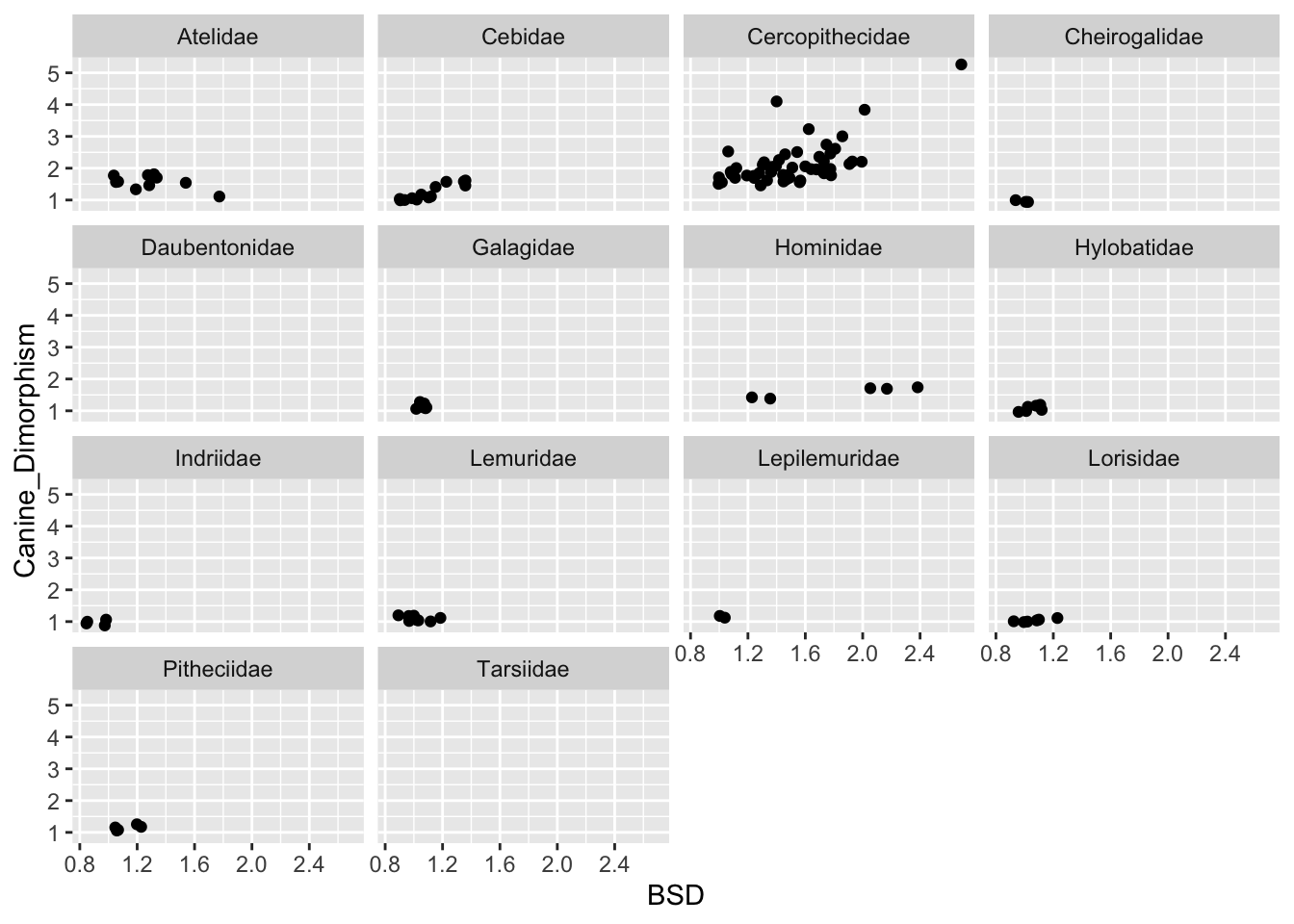
(p <- ggplot(data = d, aes(x = BSD, y = Canine_Dimorphism)) +
geom_point(na.rm = TRUE)) +
facet_wrap(~Family)
# 7
d <- d |> mutate(
"diet_strategy" = case_when(
Fruit >= 50 ~ "frugivore",
Leaves >= 50 ~ "folivore",
Fruit < 50 & Leaves < 50 ~ "omnivore",
TRUE ~ NA
)
)
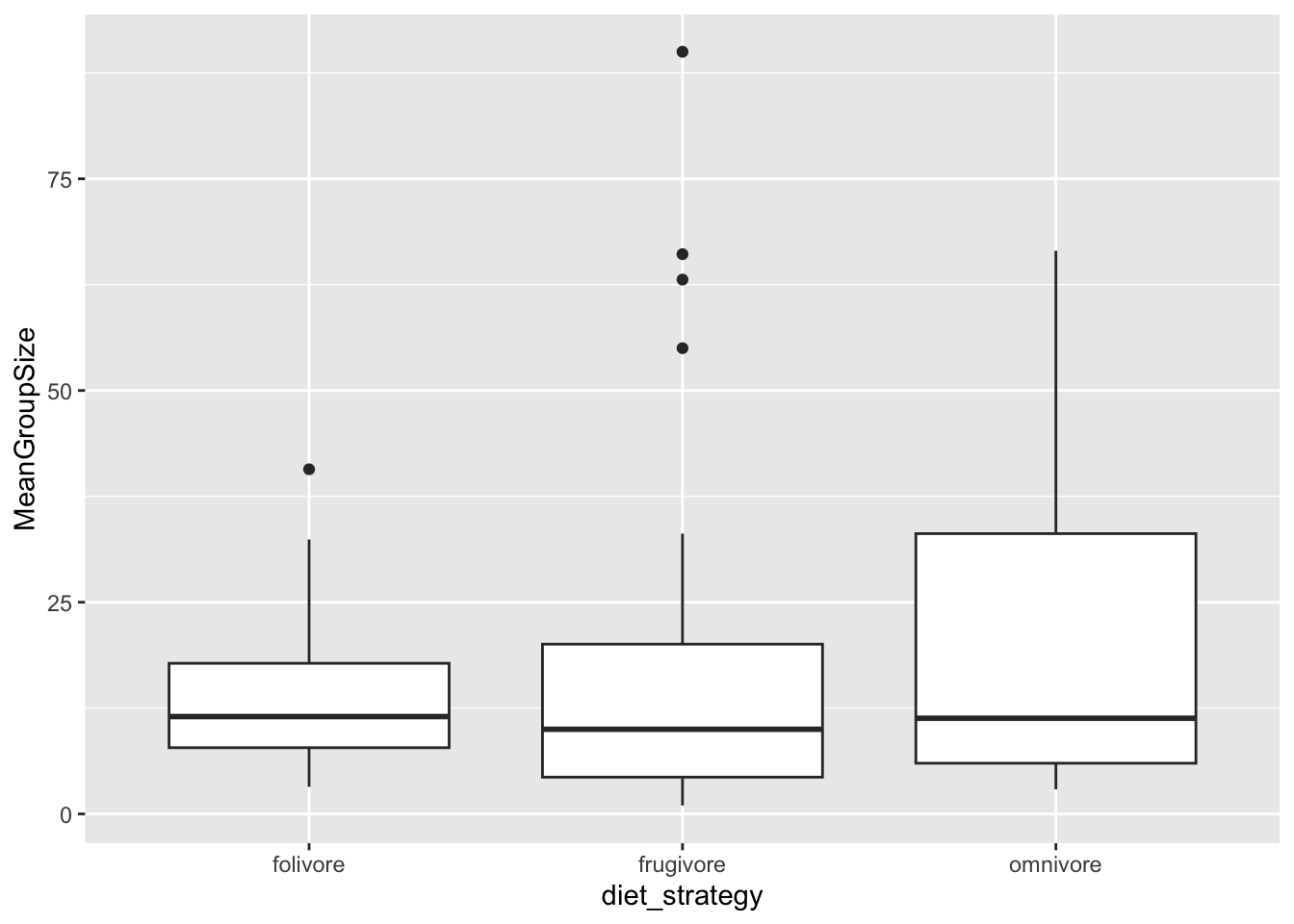
(p <- ggplot(data = filter(d, !is.na(diet_strategy)), aes(x = diet_strategy, y = MeanGroupSize)) +
geom_boxplot())
# 8
d <- d |>
mutate(Binomial = paste0(Genus, " ", Species)) |>
select(Binomial,
Family,
Brain_Size_Species_Mean,
Body_mass_male_mean) |>
group_by(Family) |>
summarise(
meanBrainSize = mean(Brain_Size_Species_Mean, na.rm = TRUE),
meanMaleBodySize = mean(Body_mass_male_mean, na.rm = TRUE)
) |>
arrange(meanBrainSize) |>
print()
```
```{r}
#| include: false
detach(package:tidyverse)
```